Collaborative Brain Wave Analysis Pipeline (Cobrawap)¶

Cobrawap is an adaptable and reusable analysis pipeline for the multi-scale, multi-methodology analysis of cortical wave activity. The pipeline ingests data from multiple measurements types of spatially organized neuronal activity, such as ECoG or calcium imaging recordings. The pipeline returns statistical measures to quantify the dynamic wave-like activity patterns found in the data.
Documentation | Publication | Introductory video | Demo on the EBRAINS Collab
Concept¶
For researchers to be able to effectively reproduce results and build on each other’s progress, it is important to not only make results openly accessible and to facilitate data sharing, but also to build the analysis workflows in a shareable and reusable manner.
Making analysis scripts available alongside results and datasets is good. What is even better is to design the analysis workflows for specific research questions in such a manner that they are general and flexible enough to be actively reused in further research. In this way, the rigor of the analysis workflow is increased, and the design of the analysis process is simplified for the researcher
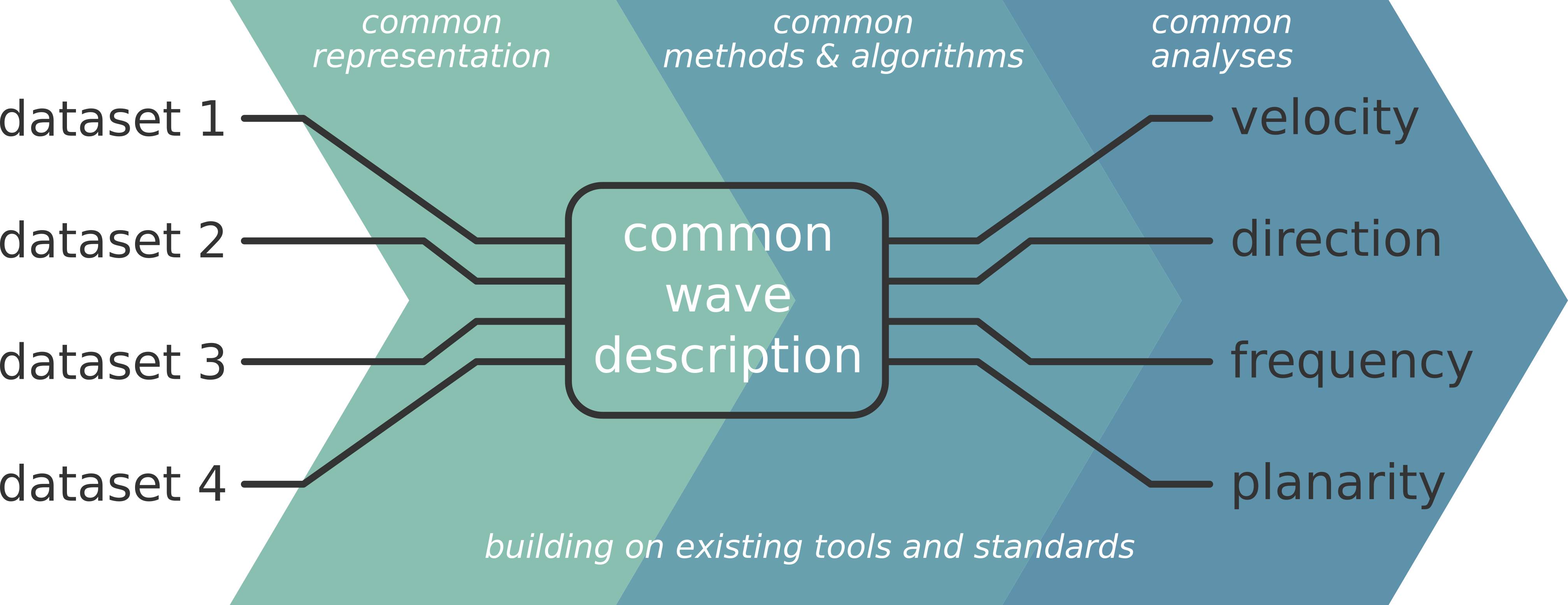
Cobrawap brings together existing analysis methods, tools, and data standards and interfaces them for the analysis and characterization of cortical wave-like activity and UP/DOWN state detections. Cobrawap serves as a space to gather the various data types exhibiting wave activity and their various analysis approaches into the same pipeline. Besides generating easily reproducible and curated results, Cobrawap facilitates the rigorous comparison between datasets of different laboratories, studies and measurement modalities. Furthermore, Cobrawap can be used in the context of model validation and benchmarking of analysis methods. Cobrawap may also act as a template for implementing analysis pipelines in other contexts.
Cobrawap features…
a user-friendly command line interface guiding the setup and usage
a hierarchical and modular pipeline framework based on the Snakemake workflow management tool
reusable method implementations (stages and blocks) for standalone applications or integration into workflows
analysis methods for electrophysiological and optical data on the characterization of cortical wave activity and local oscillations
visualization of the analysis steps and the intermediate results
intermediate results curated with annotated metadata
For further developments and feature requests refer to the Github Issues.
Citation¶
To refer to the Cobrawap software package in publications, please use:
Cobrawap (doi:10.5281/zenodo.10198748; RRID:SCR_022966)
To cite a specific version of Cobrawap please see version-specific DOIs at:
To cite Cobrawap, please use:
Gutzen, R., De Bonis, G., De Luca, C., Pastorelli, E., Capone, C., Allegra Mascaro, A. L., Resta, F., Manasanch, A., Pavone, F. S., Sanchez-Vives, M. V., Mattia, M., Grün, S., Paolucci, P. S., & Denker, M. (2022). Comparing apples to apples—Using a modular and adaptable analysis pipeline to compare slow cerebral rhythms across heterogeneous datasets. arXiv:2211.08527. https://doi.org/10.48550/arXiv.2211.08527
License¶
Cobrawap is open-source software and is licensed under the GNU General Public License v3.
Involved partners¶
Cobrawap was originally developed in the context the Human Brain Project and is part of the EBRAINS infrastructure. Partners involved in the collaboration and use case within the project (WaveScalES) are:
Forschungszentrum Jülich, Germany: Robin Gutzen, Michael Denker
Istituto Nazionale di Fisica Nucleare (INFN), Roma, Italy: Giulia De Bonis, Pier Stanislao Paolucci, Elena Pastorelli, Francesco Simula, Cristiano Capone, Chiara De Luca, Cosimo Lupo, Irene Bernava
Istituto Superiore di Sanità (ISS), Roma, Italy: Maurizio Mattia, Antonio Pazienti.
Institut d’Investigacions Biomediques August Pi i Sunyer (IDIBAPS), Barcelona, Spain: Arnau Manasanch, Miguel Dasilva, Maria V. Sanchez-Vives.
European Laboratory for Non-Linear Spectroscopy (LENS), Firenze, Italy: Anna Letizia Allegra Mascaro, Francesco Resta, Francesco Pavone.
University of Milano (UniMi), Italy: Andrea Pigorini, Thierry Nieus, Marcello Massimini
Unité de Neurosciences, Neuroinformatics Group, CNRS, France: Andrew Davison
Further Context¶
Software Ecosystem¶
The functionality offered by Cobrawap builds on existing software tools and services.
Neo improves interoperability between Python tools for analyzing, visualizing, and generating electrophysiology data by providing a common, shared data object model. The Neo data representation provides a hierarchical data and metadata description for a variety of data types including intracellular and extracellular electrophysiology electrical data with support for multi-electrode as well as optical recordings. Furthermore, it supports a wide range of neurophysiology file formats to facilitate reading data from most common recording devices.
The Electrophysiology Analysis Toolkit, Elephant, is an open-source Python library for analysis methods. It focuses on providing fast and reliable implementations for generic analysis functions for spike train data and time series recordings from electrodes. As community centered project Elephant aims to serve as a common platform for analysis codes from different laboratories, and a consistent and homogeneous analysis framework.
The Neuroscience Information Exchange, NIX, format is an API and data format to store scientific data and metadata in a combined representation. Its structure is inspired by common types of neuroscience data, and it acts as one of the primary data formats for the Neo data object model.
The WaveScalES project¶
Sleep is present in all animal species notwithstanding the risk associated with the disconnection from the environment (e.g. predation) and the reduction of time available for food search and reproduction. Indeed, it is known that the human brains need healthy sleep, as chronic sleep deprivation reduces cognitive performances. The goal of the WaveScalES collaboration of the Human Brain Project is to unveil the underlying mechanisms of deep sleep, anesthesia and coma, the emergence toward wakefulness, and the link between sleep and learning, taking advantage of cortical slow wave activity (SWA) and investigating it with experimental data, analysis tools, modulation techniques, theoretical models and simulations of such states and of the transition to wakefulness.
